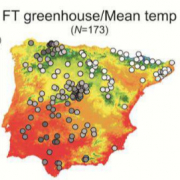
Regional GWAS of flowering adaptation
Genome-wide signatures of flowering adaptation to climate temperature: regional analyses in a highly diverse native range of Arabidopsis thaliana
Collaborative work with Carlos Alonso-Blanco and Xavi Picó
Tabas-Madrid et al.: Plant Cell Environ. published online Mar 8.
Current global change is fueling an interest to understand the genetic and molecular mechanisms of plant adaptation to climate. In particular, altered flowering time is a common strategy for escape from unfavorable climate temperature. In order to determine the genomic bases underlying flowering time adaptation to this climatic factor, we have systematically analysed a collection of 174 highly diverse A. thaliana accessions from the Iberian Peninsula. Analyses of 1.88 million SNPs provide evidence for a spatially heterogeneous contribution of demographic and adaptive processes to geographic patterns of genetic variation. Mountains appear to be allele dispersal barriers, whereas the relationship between flowering time and temperature depended on the precise temperature range. Environmental genome-wide associations (EGWA) supported an overall genome adaptation to temperature, with 9.4% of the genes showing significant associations. Furthermore, phenotypic genome-wide associations (PGWA) provided a catalogue of candidate genes underlying flowering time variation. Finally, comparison of EGWA and PGWA genomic regions identified known (TSF, FRL1 and CKB1) and new (ESM1 and VDAC5) genes as candidates for adaptation to climate temperature by altered flowering time. Thus, this regional collection provides an excellent resource to address the spatial complexity of climate adaptation in annual plants.