On bioRxiv: Link between ACD6 and NLR signaling
Modulation of ACD6 dependent hyperimmunity by natural alleles of an Arabidopsis thaliana NLR resistance gene
Zhu, W., Zaidem, M., Van de Weyer, A.-L., Gutaker, R. M., Chae, E., Kim, S.-T., Bemm, F., Li, L., Schwab, R., Unger, F., Beha, M. J., Demar, M., and Weigel, D.
bioRxiv posted April 13, 2018
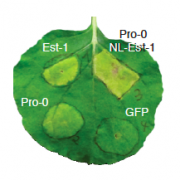
Plants defend themselves against pathogens by activating an array of immune responses. Unfortunately, immunity programs may also cause unintended collateral damage to the plant itself. The quantitative disease resistance gene ACCELERATED CELL DEATH 6 (ACD6) serves as a nexus for the trade-off between growth and pathogen resistance in wild populations of Arabidopsis thaliana. An autoimmune allele, ACD6-Est, first identified in the natural accession Est-1, is found in over 10% of wild strains, even though it causes a clear fitness penalty under optimal growth conditions. There is, however, extensive variation in the strength of the autoimmune phenotype expressed by strains with an ACD6-Est allele, indicative of genetic modifiers. Quantitative genetic analysis suggests that the population genetic basis of ACD6 modulation is complex, with different strains often carrying different large-effect modifiers. One modifier is SUPPRESSOR OF NPR1-1, CONSTITUTIVE 1 (SNC1), located in a highly polymorphic cluster of nucleotide-binding domain and leucine-rich repeat (NLR) immune receptor genes, which are prototypes for qualitative disease resistance genes. Allelic variation at SNC1 correlates with ACD6-Est activity in multiple accessions, and a common structural variant affecting the NL linker sequence can explain differences in SNC1 activity. Taken together, we find that an NLR gene can mask the activity of an ACD6 autoimmune allele in natural A. thaliana populations, thereby linking different arms of the plant immune system.